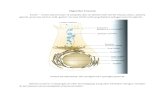
Nitrogenase enzyme complex
Transcript of Nitrogenase enzyme complex

Redrawn from www.asahi-net.or.jp/~it6i-wtnb/BNF.html
Nitrogenase enzyme complex
Physical association of nif genes in Klebsiella pneumoniae
NitrogenaseMoFe protein
Fe proteinElectron transport
AssemblingFe-Mo-Cofactor
Regulator
H D K T Y E NX U SVWZM F L A B QJ
α βγ γαβ

Redrawn from http://www.science.siu.edu/microbiology/micr425/425Notes/12-NitrFix.html
NtrC NtrC
in absence ofN-compounds
NtrB
ADP ATP
nifLA operonnitrC binding site σ54 = nitrAbinding site
nif structural genes σ54 = nitrAbinding site
nifA binding site
P
Regulation of nitrogen fixation (K. pneumoniae)

Nitrogen present, no transcription
Function of NtrA, σ54 , thenitrogen σ factor

Nitrogen absent, NtrB phosphorylates NtrC,which activates RNA polymerase
Function of NtrA, σ54 , thenitrogen σ factor
P

Redrawn from http://www.science.siu.edu/microbiology/micr425/425Notes/12-NitrFix.html
NtrC NtrC
in absence ofN-compounds
NtrB
ADP ATP
nifLA operonnitrC binding site σ54 = nitrAbinding site
nif structural genes σ54 = nitrAbinding site
nifA binding site
P
Regulation of nitrogen fixation (K. pneumoniae)

Redrawn from http://www.science.siu.edu/microbiology/micr425/425Notes/12-NitrFix.html
NifANifANifL
transcription
NtrC
in absence ofN-compounds
NtrB
ADP ATP
nifLA operonσ54 = nitrAbinding site
NtrC P
N-compound regulation of NifLA operon
nif structural genes σ54 = nitrAbinding site
nifA binding site

Redrawn from http://www.science.siu.edu/microbiology/micr425/425Notes/12-NitrFix.html
NifANifA
NifL
transcription
transcription
NtrC
in absence ofN-compounds
NtrB
ADP ATP
nifLA operonσ54 = nitrAbinding site
nif structural genes σ54 = nitrAbinding site
N-compound regulation of NifLA operon
NtrC P

Redrawn from http://www.science.siu.edu/microbiology/micr425/425Notes/12-NitrFix.html
NifANifANifANifA
NifLNifLin presence
of O2 orN-compounds
transcription
NtrC
in absence ofN-compounds
NtrB
ADP ATP
nifLA operonσ54 = nitrAbinding site
nif structural genes σ54 = nitrAbinding site
nifA binding site
Oxygen and N-compound regulation of nif structural genes via nifL
NtrC P

Measuring N2 fixation rates

Acetylene reduction assay
• Football has been filled withacetylene
• Glass jars contain the plantsamples being evaluated
• Sterile vacutainers (normallyused to collect blood) areused to take the gas samplefollowing incubation
• Several hundred samplescan be taken each day
http://www.soils.umn.edu/academics/classes/soil3612/Nitrogen_Fixation/Measurement.htm

Acetylene reduction assay
• A typical trace followinggas chromatography
• The greatest peaks areof residual acetylene
• Those next to them theethylene peak
http://www.soils.umn.edu/academics/classes/soil3612/Nitrogen_Fixation/Measurement.htm

Hydrogen evolution assay
• Reduction of dinitrogen to ammonia
N2 + 8H+ + 8e- → 2NH3 + H2
• H2 is evolved at ratio of 1 moleculeper 2 molecules of N2 reduced
– So, can use hydrogen sensor tomeasure H2 evolution to quantify N2
fixation

Hydrogen evolution assay
• Reduction of dinitrogen to ammonia
N2 + 8H+ + 8e- → 2NH3 + H2
• H2 is evolved at ratio of 1 moleculeper 2 molecules of N2 reduced
– So, can use hydrogen sensor tomeasure H2 evolution to quantify N2
fixation

Hydrogen evolution assay

The operation of nitrogenase. The iron protein (Fe) takes electronsfrom central metabolism electron carriers and transfers them to themolybdenum iron protein (MoFe) expending a fair amount of ATP.N2 is converted to ammonia and the electrons in H2 are recycled byhydrogenase. (D. Benson)

Stable isotope assays
GC Mass separationCombustion
detector
ion source
magnet

mass 29 (15N14N)
ion source
magnet
mass 30 (15N2)
mass 28(14N2)
Mass Separation

Stable isotope lab

Crop Common name δ15N
Agropyrontrichophorum
Pubescentwheatgrass
5.13
A.elongatum Tall wheatgrass 3.04
A.dasystachyum NorthernWheatgrass
3.00
Elymus angustus Wild ryegrass 2.31
Melilotus officinales Yellow sweetclover
-0.72
Medicago sativa Alfalfa 0.82
Astragalus cicer Milk vetch 1.90
Atmospheric N2 0.00
Soil N 7.00
Table 15-3 (pg. 380 of text)

Lifestyles of N2 fixing bacteria(diazotrophs)
• Free living
• Living in consortia– e.g. stromatolites, soil crusts
• Plant associative (living in rhizosphere)
• Symbiotic

Middle Proterozoic formations ofthe Hakatai Shale in Grand
Canyon National Park.Lens cap is 55 mm.
Livingstromatolites
Diazotrophicbacteria
in consortia

Diazotrophicbacteria
in consortiasoil crusts

Cyanobacteria insoil crusts

Diazotrophicbacteria
in consortia

Cyanobacteria• Photosynthetic and
dinitrogen fixing– heterocysts separate
the two functions
Anabaena
Microcystis
Nostoc
Free-living

Cyanobacteria• Oldest known fossils
– 3.5 bybp (oldest rocks are 3.8 bypb)
filamentous Palaeolyngbya
colonial chroococcalean

Cyanobacteria heterocysts
Heterocyst

Symbiotic N2-fixation: Azolla - Anabena
S. Navie

Symbiotic N2-fixation: Azolla - Anabena

Rice-Azolla-Fish, China
Azolla to feed cows, Thailand
Rice-Azolla-Ducks, KoreaTakao Furuno
Symbiotic N2-fixation: Azolla - Anabena

© Paul Cox
Cycas micronesica

Cycad root nodules

© Paul Cox
Cycas micronesicaβ-N-methylamino-L-alanine (BMAA)

Guam flying fox(Pteropus mariannus)
bio-magnification
β-N-methylamino-L-alanine (BMAA)

Flying Fox with Prunes and Cream Sauce
6 flying foxes (in case you are wondering, these are bats) 1 pound prunes 1_ cup white wine salt, pepper 1/4 cup flour 2 oz. butter 1 tbsp red currantjelly 1 cup thick cream
Remove the flesh from the flying foxes. Either plunge the animals in boilingwater for a while, then skin them and remove the flesh from the bones, orroast the animals for a little over an open fire, remove, and when cool, breakopen down the backbone and remove the flesh from the skin.
Soak the prunes overnight in 1 cup of the wine, then heat for about tenminutes in the wine before using. Season the flying fox meat with salt andpepper and roll in flour. Saute in butter over a low heat until brown. Add therest of the wine, cover and cook another 20 minutes. Add the juice from theprunes, and transfer the prunes onto a serving dish. Cook the meat in theprune juice, uncovered, for another 10 minutes, then place on the servingdish with the prunes.
The preparation of this recipe requires an ingredient which is now aprotected species.

Spinal cord
Amyotrophic Lateral Sclerosisβ-N-methylamino-L-alanine
Mimics glutamateand acts as agonistat glutamatereceptor

Frankia in alder root nodules
Actinorhizal symbioses

Frankia vesicles
Frankia root nodulesSpores &hyphae

ColletiaDicaria
Ceanothus

Ceanothus

Myrica faya
Native to Canary Islands
Actinorhizal root nodules

Invasive in Hawai’i (no native N2 - fixing pioneers)
Myrica faya

Legumes &N-fixingbacteria
© Simms
Soil-dwelling rhizobia infectlegume roots

• Signals early in infection– Complex handshaking between legume root
and rhizobium
Incorrect signal
Correctsignal

Legume & N-fixing bacteria• Rhizobia engulfed into
nodule cells
• Differentiate intobacteroids
© Simms

Illustration: M.S. Hargrove
Photo: D. Hume
Photo and illustration: R. F. Denison
Leghemoglobin

RhizobiumAgrobacterium
Sawada et al. 2003
SinorhizobiumEnsifer
Mesorhizobium,
BradyrhizobiumNitrobacter, Afipia
Methylobacterium
Sinorhizobium
DevosiaAzorhizobium
Ralstonia
Burkholderia
Rickettsia
BartonellaAminobacter, Phyllobacteriumα
β
-Rhizobial symbioses have evolved ~10 times-Nested parasites & non-symbionts
Brucella
Proteobacteria

Symbiotic plasmid of Rhizobium etli
Víctor González et al. Genome Biology 2003 4(6):R36

The nodulation genes nodABCDIJ are represented in blue
The nitrogen-fixation genes nifHDKNEXAB, fixABCX and fdxBN are represented in yellow
plasmid 42d
M. loti MAFF303099
plasmid NGR234a
M. loti MAFF303099
B. japonicum
S. meliloti pSymA
Víctor González et al. Genome Biology 2003 4(6):R36

Victor Kunin et al. Genome Res. 2005; 15: 954-959
Figure 3. Three-dimensional representation of the net of life


Hazards of symbiotic life (or an animal dispersal agent?)
Clover Root CurculioSitona hispidula (Fabricus)