Omics Data Analysis Using SOP (Search of Omics Pathway) Web … · 2009-11-17 · Omics Data...
Transcript of Omics Data Analysis Using SOP (Search of Omics Pathway) Web … · 2009-11-17 · Omics Data...
Omics Data Analysis Using SOP (Search of Omics Pathway) Web Tool
Seung Yong Lee1, Jun-Sub Kim2, Seung-Jun Kim1,2, Ji-Hoon Kim2,
[email protected], [email protected], [email protected], [email protected] Hye-Won Park1,3 , Yu Ri An1, Moon-Ju Oh2, Seung Yong Hwang1,2*
[email protected] [email protected] [email protected] [email protected]
1 Department of Biochemistry, Hanyang University, Sangrok-gu, Ansan, Gyeonggi-do, Korea 2 GenoCheck Co. Ltd., Sangrok-gu, Ansan, Gyeonggi-do, Korea
3 Department of Bionanotechnology, Hanyang University, Sangrok-gu, Ansan, Gyeonggi-do, Korea
Keywords: Database, KEGG Pathway, Omics, Toxicogenomics, Web tool 1 Introduction
Omics is a term of concept including a large-scale data in biology. Toxicogenomics is paid more attention to many researchers to investigating the genome-scale genes/proteins interaction, expression pattern and function in exposed state to specific toxicant or toxicant-related environment. The ultimate goal of toxicogenomics identifies functional context among the group of genes that are differentially or similarly coexpressed under the specific toxic substance. For these reasons, many omics databases including gene/protein annotation, function and metabolic pathway information are required to researchers, but these databases are heterogeneous and independent. 2 Method and Results
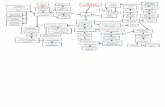
It is hard to analyze a large omics annotations and their pathway information at once. Therefore we propose SOP (Search of Omics Pathway) database system which is constructed as the integrated biological database and web tool (http://www.koreagene.co.kr/omics/sop/). The data of SOP web interface are supported as the genes/prteins annotation web table and the visualized pathway maps of KEGG PATHWAY database..
Figure 1: Schematic overview of SOP database system. Server-side DB layer was manipulated by Perl and MySQL. Server-side software layer is written in Perl for compiling user query and executing command. Client-side layer gets
P119-1
3 Discu
We believtraced to ohypothesis f Referenc [1] Aardem
“toxico [2] Lee, K
experim [3] Wan-S
KIM, SKyung
KIM., [4] Goto S
and rea [5] http://e
FiguSOPon wOrgaident
ussions
ve that SOP womics experifor a next tox
ces
ma, M.. J. ogenomics”:
K. M., Kim, Jment. Toxico
Seon LEE, GSeung-Jun Kg-Sun KANG
The intellige
S., Okuno Y.
actions in bio
en.wikipedia
ure 2: Main fr main web p
web browser. anism optiontifier or gene
will 1) appeaments, 2) lexicogenomic
& MacGreimpact of “-
J. H. & Kangol Appl Phar
Gyoum-Jung LKIM, Jun-SubG, Yong-Soon
ent data man
, Hattori M.,
ological path
a.org/wiki/-o
rame of the Spage supports
In Query Pan (human, moe express pro
ar as a usefulead to new cs experimen
gor, J. T.,-omics” tech
g, D., Designrmacol 207:2
LEE, Gyounb KIM, Seunn LEE, Ki-Se
nagement sys
, Nishioka T.
hways. Nucle
omics.
SOP web apps tree panels anel user canouse, rat) andofile). If user
l tool to perffocuses from
nts
Toxicologyhnologies. Mu
n issues in tox200-208, 200
ng-Jae LEE, Cng Yong HWAeon JEON, C
stem for toxic
., Kanehisa M
eic Acids Res
plication (Query Pane
n submit querd inputting uquery is suc
form biologicm their path
y and genetutat Res 499
xicogenomic5.
Chan-Dong YANG, Joon-SChan-Hwi UM
cogenomics.
M., LIGAND
s 30:402-404
el, Viewer Pary to SOP serser query in
ccessfully sub
cal interpretahway analys
tic toxicolog:13-25, 2002
cs using DNA
YEO, Jong-SSuk PARK, JM, Seok-Il H
J Vet Med S
D: database o
4, 2002.
anel, Annotarver by selecQuery optionbmitted
ation of genesis, and 3) s
gy in the n2.
A microarray
Soo KANG, Jae-Woong HHONG and Y
Sci 66:1335-
of chemical c
ation Panel) cting ns (a single
es or proteinssuggest new
new era of
y
Yong-Hyun HWANG, Yang-Seok
1338, 2004.
compounds
s w
f
P119-2