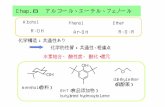
CH3 OMe CH2OHfgdfg 1 1 C:\Bruker\TopSpin3.1\examdata! S29! 13C NMR (100 MHz, CDCl 3) H3C MeO N H O...
Transcript of CH3 OMe CH2OHfgdfg 1 1 C:\Bruker\TopSpin3.1\examdata! S29! 13C NMR (100 MHz, CDCl 3) H3C MeO N H O...

S12
1H NMR (400 MHz, CDCl3)
CH3
OMe
CH2OH
4
Electronic Supplementary Material (ESI) for Organic & Biomolecular Chemistry.This journal is © The Royal Society of Chemistry 2017

S13
13C NMR (100 MHz, CDCl3)
CH3
OMe
CH2OH
4

S14
1H NMR (400 MHz, CDCl3)
CH3
OMe
CHO
5

S15
13C NMR (100 MHz, CDCl3)
CH3
OMe
CHO
5

S16
1H NMR (400 MHz, CDCl3) H3C
MeONH
HN
CH3
OMe
1a

S17
13C NMR (100 MHz, CDCl3) H3C
MeONH
HN
CH3
OMe
1a

S18
1H NMR (400 MHz, CDCl3)
H3C
MeONH
1b
NH
CH3
OMe

S19
13C NMR (100 MHz, CDCl3)
H3C
MeONH
1b
NH
CH3
OMe

S20
1H NMR (400 MHz, CDCl3) H3C
MeONH
1c
HN
CH3
OMe

S21
13C NMR (100 MHz, CDCl3) H3C
MeONH
1c
HN
CH3
OMe

S22
1H NMR (400 MHz, CDCl3)
1d
H3C
MeONH
N3
2.923.806.34
3.135.69
9.006.076.809.18
ppm12345678
Plotname: --Not assigned--

S23
1d
H3C
MeONH
N3
156.751
127.124
133.584
134.237
126.988
123.496
121.340
118.532
102.530
77.367
77.048 76.737
55.111
53.471
49.835
46.936
19.950
ppm20406080100120140160180200
Plotname: --Not assigned--

S24
1H NMR (400 MHz, CDCl3) 2a
H3C
MeONH
HN
CH3
1.023.85
4.992.00
2.983.91
5.92
ppm12345678
Plotname: --Not assigned--

S25
13C NMR (100 MHz, CDCl3)
ppm20406080100120140160180200
Plotname: --Not assigned--
2a
H3C
MeONH
HN
CH3

S26
1H NMR (400 MHz, CDCl3)
[ppm] 8 6 4 2
[rel
] 0
5
1
0 1
5 2
0 2
5
1.00
00
2.13
671.
0009
1.00
402.
0486
2.03
601.
0058
2.86
302.
4345
1.93
38
13.1
665
2.46
50
fdsfesdf 1 1 C:\Bruker\TopSpin3.1\examdata
N3O
OBn
BnO OBn
N3
8

S27
13C NMR (100 MHz, CDCl3)
ppm20406080100120140160180200
Plotname: --Not assigned--
N3O
OBn
BnO OBn
N3
8

S28
1H NMR (400 MHz, CDCl3) H3C
MeONH
O
OH
HO OH
HN
CH3
OMe
3b
[ppm] 8 6 4 2
[rel
] 0
5
1
0 1
5
2.00
00
5.54
112.
3869
2.15
63
2.26
96
6.26
04
3.15
14
1.33
91
6.36
72
1.64
88
2.04
22
1.29
681.
3528
1.79
94
1.53
471.
0759
fgdfg 1 1 C:\Bruker\TopSpin3.1\examdata

S29
13C NMR (100 MHz, CDCl3) H3C
MeONH
O
OH
HO OH
HN
CH3
OMe
3b
47.589
ppm20406080100120140160180
Plotname: --Not assigned--

S30
DNA Binding Studies
Materials: CT-DNA was purchased from CALBIOCHEM. Solutions of CT-DNA were prepared in 10mM Tris-EDTA buffer at pH 5.48 (as described in Jenkins: Jenkins, T.C. Optical Absorbance and Fluorescence Techniques for Measuring DNA-Drug Interactions. In Methods in Molecular Biology, Drug-DNA Interaction Protocols; Fox, K.R., Ed. Humana: Totowa, 1997; Vol 90, pp.195-217) and gave a 1.83:1 absorbance ratio at 260 nm and 280 nm. DNA and ligand concentrations were determined using 8452A HP Diode Array Spectrophotometer: CT-DNA, e260 = 6600 M-1 cm-1 bp-1 (1.82:1 absorbance ratio at 260 nm and 280 nm); poly(dG)•polyd(C), e253 = 7400 M-1 cm-1 bp-1; poly (dA) • poly (dT), e260 = 6000 M-1 cm-1 bp-1; poly(dA-dT)•poly(dA-dT), e262 = 6600 M-1 cm-1 bp-1; Lambda phage DNA: 5 units dissolved in 500 mL buffer provides a solution of 0.76 mM/bp. Methyl Green, e631 = 85, 300 M-1 cm-1; netropsin, e296 21,500 M-1cm-1; Hoechst 33342, e340 = 47,000 M-1 cm-1; Ethidium Bromide, e480 = 5450 M-1 cm-1; 1a-1c, 3b, e336 = 6089 M-1 cm-1; 1d, e336 = 9134 M-1 cm-1; 2a, e336 = 3045 M-1 cm-1. EB competitive experiments Constant concentrations of CT-DNA (poly(dG)•polyd(C), poly (dA)•poly (dT), poly(dA-dT)•poly(dA-dT), or Lambda phage DNA, 10 µM) and EtBr (10 µM) were titrated with increasing concentrations of the ligands (from 1 mM and 100 µM stock solutions), in the presence or absence of fixed concentrations of NaCl or the competitors methyl green or netropsin. The maximum emission wavelength was 490 nm when the excitation wavelength was 520 nm. Fluorescence titrations were recorded from 520 nm to 692 nm after an equilibration period of 3 min. Ex Slit (nm) = 10.0; Em Slit (nm) = 10.0; Scan Speed (nm/min) = 200. Viscosity Studies Viscosity experiments were performed with an Ostwald viscometer in a constant water bath at 23.0 ± 1°C. Solutions of constant DNA concentrations and varying ligand concentrations in Tris-EDTA buffer were incubated for 30 minutes. A digital stopwatch was used to record the flow time. The relative viscosity was calculated as from the following equation: 𝜂 = !!!!
!!
where t0 and t are the flow time in the absence and presence of the ligand. η is the viscosity in the presence of the ligand and η0 is the viscosity in the absence of the ligand. The data were graphed as (η/η0)1/3 vs. [ligand]/[DNA]. Circular Dichroism Studies. Small aliquots (0.6-5.0 μL) of a concentrated 1d solution (1 mM) were added to a solution (2 mL, 100 mM KCl, 10 mM SC, 0.5 mM EDTA, pH 6.8) of CT-DNA (80 μM/bp), inverted twice, and incubated for 5 min at 20 °C. The CD spectra were then recorded as an average of three scans from 220 to 310 nm and data recorded in 0.1 nm increments with an averaging time of 2 s.
1H(400 MHz) and 13C NMR (100 MHz) spectra

S31
Compound 1a Ethidium displacement studies; Kapp=2.1x106M-‐1 x 10/C50
Titration of CT DNA (10 µM) and ethidium (10 µM) with 1a: 0.005, 0.01, 0.02, 0.04, 0.05, 0.1, 0.2, 0.33, 0.45, 0.56, 0.63, 0.71, 0.83, 1.00, 1.25, 1.67,
2.50, 3.30, 5.0, 10.0, 20.0, 30.0, 40.0, 50.0, 60.0, 100.0, 200.0 µM
Trial 1: Kapp= 6.17 x 106 M-1; Trial 2: Kapp= 8.40 x 106 M-1 ; Trial 3: Kapp= 7.00 x 106 M-1
Average Kapp=7.19 ± 0.92 x 106 M-‐1
Average data for Trials 1-3
Nor
mal
ized
Flu
ores
cen
ce
-0.2
0
0.2
0.4
0.6
0.8
1
1.2
-0.2
0
0.2
0.4
0.6
0.8
1
1.2
log([1a])-9 -8 -7 -6 -5 -4 -3
-9 -8 -7 -6 -5 -4 -3
C(50)=3.09±0.02x10-6M
Plot of Δ Fluorescence vs. rbd for 1a
Δ Fl
uor
esce
nce
-0.08
-0.07
-0.06
-0.05
-0.04
-0.03
-0.02
-0.01
0
0.01
-0.08
-0.07
-0.06
-0.05
-0.04
-0.03
-0.02
-0.01
0
0.01
rbd0 5 10 15 20 25 30 35
0 5 10 15 20 25 30 35
lines intersect at rbd=4.2
[1a]=0.005 µM
[1a]=200 µM
Titration of CTDNA•ethidium with 1a
Nor
mal
ized
Flu
ores
cen
ce
-0.2
0
0.2
0.4
0.6
0.8
1
1.2
-0.2
0
0.2
0.4
0.6
0.8
1
1.2
log([1a])-9 -8 -7 -6 -5 -4
-9 -8 -7 -6 -5 -4
0mM NaCl, C(50)=3.23x10-6M10mM NaCl, C(50)=8.70x10-6M100mM NaCl, C(50)=12.6x10-6M

S32
Effect of increasing amount of EtBr (▲) and 1a (●) on relative viscosity of CT-‐DNA. EB (▲): [CT-‐DNA] = 300 µM, [EB]= 4, 26, 70, 113, 160, and 200 µM; 1a (●): [CT-‐DNA] = 300 µM, [1a]: 8, 52, 140, 226, 320, and 400 µM.

S33
Compound 1b Ethidium displacement studies; Kapp=2.1x106M-‐1 x 10/C50
Titration of CT DNA (10 µM) and ethidium (10 µM) with 1b: 0.005, 0.01, 0.02, 0.04, 0.05, 0.1, 0.2, 0.33, 0.45, 0.56, 0.63, 0.71, 0.83, 1.00, 1.25, 1.67, 2.50, 3.30, 5.0, 10.0, 20.0, 30.0, 40.0, 60.0, 80.0, 100.0 µM
Trial 1: Kapp=2.35 x 106M-1 ; Trial 2: Kapp=3.31 x 106M-1; Trial 3: Kapp=3.80 x 106M-1
3 trials: Average Kapp=3.15(±0.60)x106M-1
Plot of Δ Fluorescence vs rbd for 1b
Δ Fl
uor
esce
nce
-0.14
-0.12
-0.1
-0.08
-0.06
-0.04
-0.02
0
-0.14
-0.12
-0.1
-0.08
-0.06
-0.04
-0.02
0
rbd0 5 10 15 20 25 30 35
0 5 10 15 20 25 30 35
lines intersect at rbd=6.2
Titlration of CTDNA•ethidium with 1b
Nor
mal
ized
Flu
ores
cen
ce
-0.2
0
0.2
0.4
0.6
0.8
1
1.2
-0.2
0
0.2
0.4
0.6
0.8
1
1.2
log([1b])-9 -8 -7 -6 -5 -4 -3
-9 -8 -7 -6 -5 -4 -3
C(50)=5.89x10-6M
[1b]=0.005 µM
[1b]=100 µM

S34
Effect of increasing amount of EtBr (▲) and 1b (●) on relative viscosity of CT-‐DNA. EtBr (▲): [CT-‐DNA] = 301 µM, [EtBr]= 1.06, 2.12, 3.71, 9.54, 18.02, and 25.97 µM; 1b (●): [CT-‐DNA] = 298 µM, [1b]: 1.08, 2.17, 3.79, 9.76, 17.88, and 26.02 µM

S35
Compound 1c Ethidium displacement studies; Kapp=2.1x106M-‐1 x 10/C50
Titration of CT DNA (10 µM) and ethidium (10 µM) with 1c: 0.005, 0.01, 0.02, 0.04, 0.05, 0.1, 0.2, 0.33, 0.45, 0.56, 0.63, 0.71, 0.83, 1.00, 1.25, 1.67, 2.50, 3.30, 5.0, 10.0, 20.0, 30.0, 40.0 µM
Trial 1: Kapp=4.64 x 106M-1; Trial 2: Kapp=4.71 x 106M-1; Trial 3: Kapp=5.65 x106M-1
3 trials: Average: Kapp=5.00 (±0.46) x 106M-1
Titration of CTDNA•ethidium with 1c
Nor
mal
ized
Flu
ores
cen
ce
-0.2
0
0.2
0.4
0.6
0.8
1
1.2
-0.2
0
0.2
0.4
0.6
0.8
1
1.2
log([1c])-8.5 -8 -7.5 -7 -6.5 -6 -5.5 -5 -4.5 -4
-8.5 -8 -7.5 -7 -6.5 -6 -5.5 -5 -4.5 -4
C(50)=4.79x10-6M
Plot of Δ Fluorescence vs. rbd for 1c
Δ Fl
uor
esce
nce
-0.14
-0.12
-0.1
-0.08
-0.06
-0.04
-0.02
-0.14
-0.12
-0.1
-0.08
-0.06
-0.04
-0.02
rbd0 5 10 15 20 25 30 35
0 5 10 15 20 25 30 35
lines intersect at rbd=7.0
[1c]=0.005 µM
[1c]=40 µM

S36
Effect of increasing amount of EtBr (▲) and 1c (●) on relative viscosity of CT-‐DNA. EB (▲): [CT-‐DNA] = 300 µM, [EB]= 4, 26, 70, 113, 160, and 200 µM; 1c (●): [CT-‐DNA] = 300 µM, [1c]: 1, 2, 4, 10, 18, and 26 µM.

S37
Compound 1d Ethidium displacement studies; Kapp=2.1x106M-‐1 x 10/C50
Titration of CT DNA (10 µM) and ethidium (10 µM) with 1d: 0.005, 0.01, 0.02, 0.04, 0.05, 0.1, 0.2, 0.33, 0.45, 0.56, 0.63, 0.71, 0.83, 1.00, 1.25, 1.67, 2.50, 3.30, 5.0, 10.0, 20.0, 30.0, 40.0, 50.0, 60.0, 100.0, 200.0 µM
Trial 1 Kapp=26.45 x 106 M-1; Trial 2 Kapp=24.11 x 106 M-1; Trial 3 Kapp=22.00 x 106 M-1
3 trials: Average Kapp=24.18 (± 1.5) x 106 M-‐1
Plot of Δ Fluoresence vs. rbd for 1d
Δ Fl
uor
esce
nce
-0.14
-0.12
-0.1
-0.08
-0.06
-0.04
-0.02
0
-0.14
-0.12
-0.1
-0.08
-0.06
-0.04
-0.02
0
rbd0 5 10 15 20 25 30 35
0 5 10 15 20 25 30 35
lines intersect at rbd=9.1
[1d]=0.005 µM
[1d]=200 µM
Titration of CTDNA•ethidium with 1d
Ave
rag
e N
orm
aliz
ed F
luor
esce
nce
-0.2
0
0.2
0.4
0.6
0.8
1
1.2
-0.2
0
0.2
0.4
0.6
0.8
1
1.2
log([1d])-8.5 -8 -7.5 -7 -6.5 -6 -5.5 -5 -4.5
-8.5 -8 -7.5 -7 -6.5 -6 -5.5 -5 -4.5
C(50)=8.71x10-7M

S38
UV spectra of 1d (25µM in 10mM Tris-‐EDTA, pH=5.48) in the presence of varying concentrations of CT-‐DNA: 0, 0.25, 0.5, 1.0, 2.0, 4.0, 5.0, 10.0, 15.0, 20.0, 25.0, 30.0, 40.0, 50.0, 100 µM.
==============================================================================================
==============================================================================================
Method file : <untitled>Information : Default MethodData File : <untitled>
Overlaid Spectra:
Wavelength (nm)200 300 400 500 600 700
Abso
rban
ce (A
U)
0
0.25
0.5
0.75
1
1.25
1.5
1.75
# Name Abs<335nm> Abs<260nm>------------------------------------------------ 1 0 0.19519 0.28519 2 0.25 0.26395 0.38040 3 0.50 0.27064 0.37132 4 1.00 0.26714 0.35623 5 2.00 0.26584 0.35866 6 4.00 0.26093 0.36414 7 5.00 0.25913 0.36742 8 10.00 0.26525 0.41658 9 15.00 0.27602 0.4723210 20.00 0.28812 0.5184211 25.00 0.30500 0.5761912 30.00 0.32296 0.6245413 40.00 0.31963 0.6705814 50.00 0.31213 0.7060215 0.28162 0.86610
Report generated by : Admin Signature: ................
---------------------------------------------------------------------------------------------- *** End Fixed Wavelength Report ***----------------------------------------------------------------------------------------------
Fixed Wavelength Report Date 3/23/2017 Time 17:39:47 Page 1 of 1
[CT DNA]=100 µM
[CT DNA]=0 µM
[CT DNA]=0 µM
[CT DNA]=100 µM

S39
Effect of increasing amounts of 1d (●), ethidium bromide ( ), and netropsin ( ) on the relative viscosity of CT-DNA. R= [DNA(bp)]/[ligand]; 1d (●): [CT-‐DNA] = 300 µM, [1d]: 1, 2, 4, 10, 18, and 26 µM; ethidium bromide ( ): [CT-‐DNA] = 300 µM, [ethidium bromide]= 4, 26, 70, 113, 160, and 200 µM; netropsin ( ): [CT-‐DNA] = 300 µM, [netropsin]= 4, 26, 70, 113, 160, and 200 µM.
Viscosity data comparison for 1d, ethidium, and netropsin
(η/η
o)1/3
0.5
1
1.5
2
2.5
3
0.5
1
1.5
2
2.5
3
1/R0 0.1 0.2 0.3 0.4 0.5 0.6 0.7
0 0.1 0.2 0.3 0.4 0.5 0.6 0.7
1dethidium bromidenetropsin

S40
Affinity of 1d for M. Luteus DNA
Trial 1 Kapp=7.29x106M-1 ;
Trial 2 Kapp=9.50x106M-1 ;
Trial 3 Kapp=6.00x106M-1;
Average Kapp=7.36(±1.40)x106M-1
Affinity of 1d for poly(dA•dT)2
Trial 1 Kapp=1.71 x107M-1;
Trial 2 Kapp=2.00 x 107M-1;
Trial 3 Kapp=1.60 x 107M-1
Average Kapp=17.7±1.5 x 106M-1
Affinity of 1d for polydG•polydC
Trial 1: Kapp=18.9x106M-1
Trial 2: Kapp=18.8x106M-1
Trial 3: Kapp=17.5x106M-1
Average Kapp=18.4±0.64x106M-1
Titration of polydG•polydC•ethidium with 1d
Ave
rag
e N
orm
aliz
ed F
luor
esce
nce
0.12
0.13
0.14
0.15
0.16
0.17
0.18
0.19
0.2
0.12
0.13
0.14
0.15
0.16
0.17
0.18
0.19
0.2
log([1d])-8.5 -8 -7.5 -7 -6.5 -6 -5.5 -5 -4.5 -4
-8.5 -8 -7.5 -7 -6.5 -6 -5.5 -5 -4.5 -4
C(50)=1.15x10-6M
Titration of (polydA•dT)2•ethidium with 1d
Nor
mal
ized
Flu
ores
cen
ce
-0.2
0
0.2
0.4
0.6
0.8
1
1.2
-0.2
0
0.2
0.4
0.6
0.8
1
1.2
log([1d])-9 -8 -7 -6 -5 -4
-9 -8 -7 -6 -5 -4
C(50)=1.17x10-6M
Titration of MLuteus DNA•ethidium with 1d
Nor
mal
ized
Flu
ores
cen
ce
0
0.2
0.4
0.6
0.8
1
1.2
0
0.2
0.4
0.6
0.8
1
1.2
log([1d])-8.5 -8 -7.5 -7 -6.5 -6 -5.5 -5 -4.5 -4
-8.5 -8 -7.5 -7 -6.5 -6 -5.5 -5 -4.5 -4
C(50)=2.88x10-6M

S41
Affinity of 1d for Lambda Phage DNA
Trial 1: Kapp=9.05x106M-1
Trial 2: Kapp=8.05x106M-1
Trial 3: Kapp=6.95x106M-1
Average Kapp=8.02±0.71x106M-1
Affinity of 1d for polydA•polydT
Trial 1: Kapp=20.2x106M-1
Trial 2: Kapp=16.0x106M-1
Trial 3: Kapp=23.4x106M-1
Average Kapp=20.0±2.5x106M-1
Affinity of netropsin for lambda phage DNA
Trial 1: Kapp=4.18x105M-1
Trial 2: Kapp=3.57x105M-1
Trial 3: Kapp=3.63x105M-1
Average Kapp=3.79±0.26x105M-1
Titration of Lambda Phage DNA•ethidum with 1d
Nor
mal
ized
Flu
ores
cen
ce
-0.2
0
0.2
0.4
0.6
0.8
1
1.2
-0.2
0
0.2
0.4
0.6
0.8
1
1.2
log([1d])-9 -8 -7 -6 -5 -4
-9 -8 -7 -6 -5 -4
C(50)=2.53x10-6M
Titration of polydA•polydT•ethidium with 1d
Nor
mal
ized
Flu
ores
cen
ce
-0.2
0
0.2
0.4
0.6
0.8
1
1.2
-0.2
0
0.2
0.4
0.6
0.8
1
1.2
log([1d])-9 -8 -7 -6 -5 -4
-9 -8 -7 -6 -5 -4
C(50)=1.06x10-6M
Titration of Lambda Phage DNA•ethidium with netropsin
Nor
mal
ized
flu
ores
cen
ce
-0.2
0
0.2
0.4
0.6
0.8
1
1.2
-0.2
0
0.2
0.4
0.6
0.8
1
1.2
log([netropsin])-9 -8 -7 -6 -5 -4 -3 -2
-9 -8 -7 -6 -5 -4 -3 -2
C(50)=5.95x10-5M

S42
Affinity of netropsin for CT DNA
Trial 1: Kapp=3.30x105M-1
Trial 2: Kapp=1.80x105M-1
Trial 3: Kapp=1.24x105M-1
Average Kapp=2.11±0.79x105M-1
No competitor, Ka=2.47±1.5 x107 M-‐1; 30µM NP, Ka=1.40±1.2 x107 M-‐1; 30µM MG, 1.05±0.9 x 10-‐7 M-‐1
Titration of CTDNA•ethidium with 1d
Nor
mal
ized
Flu
ores
cen
ce
-0.2
0
0.2
0.4
0.6
0.8
1
1.2
-0.2
0
0.2
0.4
0.6
0.8
1
1.2
log([1d])-9 -8 -7 -6 -5 -4
-9 -8 -7 -6 -5 -4
no competitor, C(50)=0.85 μM 30 μM NP, C(50)= 1.5 μM 30 μM MG, C(50)= 2.0 μM
Titration of CTDNA•ethidium with netropsin
Nor
mal
ized
flu
ores
cen
ce
0.3
0.4
0.5
0.6
0.7
0.8
0.9
1
1.1
0.3
0.4
0.5
0.6
0.7
0.8
0.9
1
1.1
log([netropsin])-9 -8 -7 -6 -5 -4 -3
-9 -8 -7 -6 -5 -4 -3
C(50)=8.6x10-5M

S43
Figure 5. The 220-310 nm region of the CD spectrum of solutions of CT DNA (80 µM) in the absence (black line) and presence of various concentrations of 1d: blue line, 0.15 µM 1d; light green line, 0.30 µM 1d; orange line, 0.45 µM 1d; red line, 1.1 µM 1d; dark green line 2.00 µM 1d
Wavelength (nm)
CD
(mde
g)
!
!!
!! 0.00 µM 1d + 80 µM CTDNA0.15 µM 1d + 80 µM CTDNA0.30 µM 1d + 80 µM CTDNA0.45 µM 1d + 80 µM CTDNA1.10 µM 1d + 80 µM CTDNA2.00 µM 1d + 80 µM CTDNA

S44
10mM NaCl: Kapp=2.25 x 107M-‐1 100mM NaCl: Kapp=2.25 x 107M-‐1 1M NaCl: Kapp=5.30 x 106M-‐1
Titration of CTDNA•ethidium with 1d, 10 mM NaClN
orm
aliz
ed F
luor
esce
nce
0
0.2
0.4
0.6
0.8
1
1.2
0
0.2
0.4
0.6
0.8
1
1.2
log([1d])-8.5 -8 -7.5 -7 -6.5 -6 -5.5 -5 -4.5
-8.5 -8 -7.5 -7 -6.5 -6 -5.5 -5 -4.5
C(50)=9.33 x10-7M
Titration of CTDNA•ethidium with 1d, 100 mM NaCl
Nor
mal
ized
FLu
ores
cen
ce
0
0.2
0.4
0.6
0.8
1
1.2
0
0.2
0.4
0.6
0.8
1
1.2
log([1d])-8.5 -8 -7.5 -7 -6.5 -6 -5.5 -5 -4.5
-8.5 -8 -7.5 -7 -6.5 -6 -5.5 -5 -4.5
C(50)=9.33x10-7M
Titration of CTDNA•ethidium with 1d, 1M NaCl
Nor
mal
ized
Flu
ores
cen
ce
-0.2
0
0.2
0.4
0.6
0.8
1
-0.2
0
0.2
0.4
0.6
0.8
1
log([1d])-8.5 -8 -7.5 -7 -6.5 -6 -5.5 -5 -4.5 -4
-8.5 -8 -7.5 -7 -6.5 -6 -5.5 -5 -4.5 -4
C(50)=3.98x10-6M
Titration of CTDNA•ethidium with 1d
Nor
mal
ized
flu
ores
cen
ce
-0.2
0
0.2
0.4
0.6
0.8
1
1.2
-0.2
0
0.2
0.4
0.6
0.8
1
1.2
log([1d])-9 -8 -7 -6 -5 -4
-9 -8 -7 -6 -5 -4
10mM NaCl, C(50)=0.95x10-6M100mM NaCl, C(50)=0.97x10-6M1M NaCl, C(50)=3.72x10-6M

S45
Model for bis-‐intercalation of ligand 1d in the major groove of DNA sequence 5’-‐ATGCAT-‐3’,
generated by Autodock Vina using DNA (PDB 1x95) and 1d minimized by Spartan 14 for Macintosh

S46
Additional binding modes for the association of 1d with DNA sequence 5’-‐ATGCAT-‐3’,
generated by Autodock Vina using DNA (PDB 1x95) and 1d minimized by Spartan 14 for Macintosh

S47
Compound 2a Ethidium displacement studies; Kapp=2.1x106M-‐1 x 5/C50
Titration of CT DNA (10 µM) and ethidium (10 µM) with 3b: 0.005, 0.01, 0.02, 0.04, 0.05, 0.1, 0.2, 0.33, 0.45, 0.56, 0.63, 0.71, 0.83, 1.00, 1.25, 1.67, 2.50, 3.30, 5.0, 10.0, 20.0, 30.0, 40.0, 50.0, 60.0, 100.0, 200.0 µM
Trial 1: Kapp= 4.36 x105M-1; Trial 2: Kapp=5.0x105M-1; Trial 3: Kapp= 4.79x105M-1
3 trials: Average Kapp= 4.72±0.27x105M-‐1
Stoichiometry Analysis for 2a
ΔFluorescence
-0.05
-0.04
-0.03
-0.02
-0.01
0
-0.05
-0.04
-0.03
-0.02
-0.01
0
rbd0 5 10 15 20 25 30 35
0 5 10 15 20 25 30 35
Lines intersect at rbd=4.02
Titration of CTDNA•ethidium with 2a
Nor
mal
ized
Flu
ores
cen
ce
-0.2
0
0.2
0.4
0.6
0.8
1
1.2
-0.2
0
0.2
0.4
0.6
0.8
1
1.2
log([2a])-9 -8 -7 -6 -5 -4 -3
-9 -8 -7 -6 -5 -4 -3
C(50)=3.72x10-5M
[2a]=0.005 µM
[2a]=200 µM

S48
Effect of increasing amount of EtBr (▲) and 2a (●) on relative viscosity of CT-‐DNA. EtBr (▲): [CT-‐DNA] = 300 µM, [EtBr]= 4, 26, 70, 113, 160, and 200 µM; 2a (●): [CT-‐DNA] = 300 µM, [2a]: 1, 2, 4, 10, 18, and 26 µM.

S49
Compound 3b Ethidium displacement studies; Kapp=2.1x106M-‐1 x 10/C50
Titration of CT DNA (10 µM) and ethidium (10 µM) with 3b: 0.005, 0.01, 0.02, 0.04, 0.05, 0.1, 0.2, 0.33, 0.45, 0.56, 0.63, 0.71, 0.83, 1.00, 1.25, 1.67, 2.50, 3.30, 5.0, 10.0, 20.0, 50.0 µM
Trial 1: Kapp=6.88 x106M-1; Trial 2: Kapp=6.99 x106M-1; Trial 3: Kapp=5.55 x106M-1
3 trials: Average Kapp=6.47±0.65 x106M-1
Plot of Δ Fluorescence vs rbd for 3b
Δ Fl
uor
esce
nce
-0.12
-0.1
-0.08
-0.06
-0.04
-0.02
0
-0.12
-0.1
-0.08
-0.06
-0.04
-0.02
0
rbd0 10 20 30 40 50 60
0 10 20 30 40 50 60
lines intersect at rbd=10
Titration of CTDNA•ethidium with 3b
Nor
mal
ized
Flu
ores
cen
ce
-0.2
0
0.2
0.4
0.6
0.8
1
1.2
-0.2
0
0.2
0.4
0.6
0.8
1
1.2
log(3b])-8.5 -8 -7.5 -7 -6.5 -6 -5.5 -5 -4.5
-8.5 -8 -7.5 -7 -6.5 -6 -5.5 -5 -4.5
C(50)=1.81±01x10-6M
[3b]=0.005 µM
[3b]=50 µM

S50
Effect of increasing amount of EtBr (▲) and 3b (●) on relative viscosity of CT-‐DNA. EtBr (▲): [CT-‐DNA] = 300 µM, [EtBr]= 4, 26, 70, 113, 160, and 200 µM; 3b (●): [CT-‐DNA] = 300 µM, [3b]: 1, 2, 4, 10, 18, and 26 µM.